Archive : Article / Volume 2, Issue 1
- Research Article | DOI:
- https://doi.org/10.58489/2836-5003/008
Transmitting, Mutational and Pandemic Nature of Corona: The Role of Transposons, IS1: Tn: IS1 and Crispr- Cas-9
1Ex Faculty IIT-Kharagpur.721302, India, IGE (Institute of Genetic Engineering) Badu- Kolkata, India
Nitosh Kumar Brahma, Ex Faculty IIT-Kharagpur.721302, India, IGE (Institute of Genetic Engineering) Badu- Kolkata, India.
Nitosh Kumar Brahma. Transmitting, Mutational and Pandemic Nature of Corona: The Role of Transposons, IS1: Tn: IS1 and Crispr- Cas-9. Arch. Immunol. Res. Ther. Vol. 2 Iss. 1. (2023) DOI: 10.58489/2836-5003/008
© 2023 Nitosh Kumar Brahma, this is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
- Received Date: 18-01-2023
- Accepted Date: 27-01-2023
- Published Date: 30-01-2023
Transposons, Alveoli, Transcription, Translation, Mutation, and Infections.
Abstract
Corona/ Covid19/Omicron / SARS/ MARS similar viruses have no individual genetic mechanism to make them encapsulated spore-forming microbes similar to bacteria, fungi, and algal blooms involved in spreading allergy. Transmission in all such cases needs fluid dynamics of water and air. This flow does not require any support of matrix, i.e., nanoparticles of carbon, silicates, and dust of animate inanimate molecules. They enter into lung alveoli surface epithelial cells, adhere on the surface of body saliva, and originated due to cold and other infections. In support of ACE-I and II (Angiotensin Converting Enzymes I and II). So, there are two release mechanisms are existing in corona infection; one is the release of coronavirus from the alveoli cells with matrix binds of one co-morbid patient directly to other healthy recipients at the distance of 6 m in a closed environment by its self-development irritation and allergy, as received by sudden irritating smoke developed from the kitchen, pollen, that grows during grass cutting, cleaning, andsanitization. Transcription of m-RNA, in a matrix-based single corona, after entering into lung alveoli starts to transcribe and translate to many molecules of coronaviruses, damages alveoli cells of the lung, caused due by individual physiological conditions, old (above 70 years) and young (below 40), co- morbid(sick) and healthy people at similar age groups. IS: Tn: IS), CRISPR/Cas-9 DNA transposons existing in corona m-RNA starts to act during transcription and translation.
Introduction
An AAIR( Ant adherent Immune Response) experimental was initiated in Bulb/C mic against serotype 026: EPEC (Enteropathogenic (invasive) Escherichia coli) a fatal diarrheagenic E.coli, killing mice overnight at ( 8 hours) found unique support of hybrid genetically engineered E.coli, carrying MRHU(+) donor plasmid in support of trascojugated pRT ( Plasmid Ronald Taylor)of Yale University, USA, 1980, expressed MRHU(+) plasmid in auxotrophic genetically Engineered (GE) Immune responses, revealed that the mice were resistant against the donors 026:EPEC, even at the highest dose of 10^8 cells / 0.2 ml, intraperitoneal inoculum, without pre-inoculated Balb/ C mice were killed at a low intraperitoneal dose of 101 cells / 0.2 ml. The molecular investigation revealed that some IS-based transposons support the expression of MRHU (+) hemagglutination in hybrid genes of Escherichia coli. By increasing antibiotic profiles, as occurred due to IS (Insertion Sequence) as represented by IS1, IS2, IS3, -.IS10 DNA sequence changes the flanking nature of transposons (Tn) “IS: Tn:IS” through antibiotic resistance profile initiate illegitimate recombination and was carefully studied in 1980 in the lab of Prof. H. Saedler, Biology-III Genetic Engineering, ALU, Freiburg, Germany. In case of increasing fatality of the corona, antibiotic treatments is recommended.
Considering the fatality rate of corona as pandemic globally, the author has attempted to recapitulate the possible application of AAIR, IS:Tn:IS, and CRISPR/Cas-9 in designing a vaccine against corona, to prevent coronavirus to adhere on trachea and lung cells, to repair and bring healthy life to mankind. IS: Tn:IS was observed/studied/discovered in Maize by Barbara Mc Clintock, USA, Peter Starlinger and Heinz Saedler, Germany studied the same in bacteria. Due to spontaneous mutations in the corona, it is speculated by the author, that “IS: Tn:IS” DNA sequences might be present in the corona, to mutate and to change the adhering, invasive, and infective spike protein. The presence of the IS-Tn-IS sequence has not been studied in SARS-COV-2, COVID-19, and Corona. The sequence length, varied from IS1 (0.8kb) to IS10, (1.2 kb) (kilobase pairs) showed the potentiality spontaneous to do illegitimate recombination. If any of the sequence IS1, IS2, IS3…IS10 found common among HIV, MERS, SARS, Influenza, and H1N1 viruses, it could be useful to develop AAIR. Both the AAIR vaccine concept and the recent revolution in DNA editing, Crispr/cas-9(Clustered, Repeat Interspaced Short Palindromic Repeat), / (Crispr- associated protein-9) could essentially be used, bidirectional in corona vaccine to protect against pandemics and their repeats in future.
The IR mechanisms which are proposed are supposed to be controlled by IS: Tn:IS transposons failed to attach the spike proteins. Hybrid DNA as involved in the formation of AAIR as a monoclonal antibody will block the adherence of corona spikes. Crispr/cas-9 will repair in addition to the transmitting, invasive nature of the corona. In the process the programmable DNA of non-pathogenic influenza virus. The author describes a few of those possible mechanisms of CRISPR transposons involved in changing protein structures of the spike protein of the corona. Jumping genes, transposons, if programmed non-pathogen m-RNA could be used in the form of a vaccine. Bacteriophage Lambda, x similar and other fusion to change the mode of infection to coccucea, are the present thought, changing the direction of the corona to human trillion microbes involved in Pneumonia.
Materials and Methods
The study inspired not only to study genetic rearrangements and illegitimate recombination, as occurred when corona infections are moving downwards (i.e., many varieties of antibiotics are applied, various sanitations are used in commonplaces, when for survivals corona stats to mutate and exposed as antibiotic-resistant human system surviving with bacterial strains, namely pneumococcal strains significantly participate the microevolution of the living world, Fig. 1 (D), and shows the segregating color changes among maize seeds, as caused by transposons, (E) is reproduced with the legendary personality of Barbara McClintock, who identified color changes in maize seeds due to the presence of spm transposons in maize seeds DNA in 1931 and received the Nobel Prize in 1983, [1].
IS-based Tn (Transposons) is involved in higher and lower groups of plant and animal kingdoms, including the evolution of viruses and bacteria. The camouflaging infective nature of SARS COV-2 coronavirus, covid-19 could be associated with Tn, as proposed by the author, [10-14]. Jumping genes as retrotransposons [9] hijack special cells called nurse cells, producing the invasive nature of DNA driving evolution, and causing diseases.
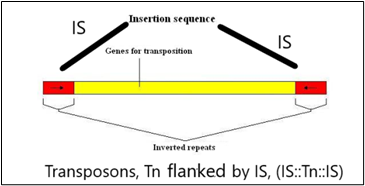
Figure1A:
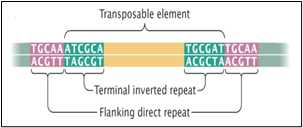
Figure1B:
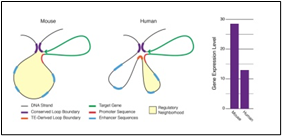
Figure1C:

Figure1D:

Figure1E:
Figure 1A, shows one transposon, flanked by IS elements (IS::Tn::IS), Figure 1B, shows the repeats and inverted repeats Dof NA sequences involved in IS elements, Figure 1C, shows the mechanism of Tn enzymes involved in illegitimate recombination and rearrangements. “Cut and Paste and Copy and Paste “foreign DNA has been identified in such illegitimate recombination where IS elements support to insert foreign DNA into host recipient DNA [1], Figure. C-2, Histogram compares the chromatin folding frequencies among mouse and human. It shows that transposons-based chromatin folding in mice is higher compared to a human. IR(Immune Response) in mice is stable, compared to humans MHC-HLA gene, responsible for IR in humans chromosome 17 is susceptible compared to chromosome 6 in mice [2-9] Chromatin loops are important for gene regulation because define a gene's regulatory neighborhood, which contains the promoter and enhancer sequences responsible for determining its expression level. Remarkably, transposable elements (TEs) are responsible for creating around 1/3 of all loop boundaries in the human and mouse genomes and contribute up to 75% of loops unique to either species. When a TE creates a human-specific or mouse-specific loop it can change a gene's regulatory neighborhood, leading to altered gene expression. The illustration shows a hypothetical region of the human and mouse genomes in which four enhancer sequences for the same target gene fall within a conserved loop. In this example, a TE-derived loop boundary in the human genome (orange bar) shrinks the regulatory neighborhood, preventing two of four enhancers from interacting with their target gene's promoter sequence. The net result is reduced gene expression in humans relative to mice. Looping variations such as these appear to be an important underlying cause of differential gene regulation across species and between different human cell types, suggesting that TE activity may play significant roles in evolution and disease. The method was studied by radio isotopic IS DNA labeling and southern hybridization. The method proves also how one IS elements jump from the plasmid to the chromosome and back to the plasmid. Almost half of our DNA sequences are made up of jumping genes, known as transposons. They jump around the genome in developing sperm and egg cells and are important in cellular evolution [15] and cause new mutations that lead to diseases. Remarkably little is known about when and where these movements started in the development of reproductive cells. The key process ensures their propagation for future generations with possible genetic disorders. Animals have developed a powerful use suppressing activities of jumping genes, non-coding pi RNAs [15-17], that recognize jumping genes and suppress their activity.
Some information has been recorded that the COVID-19 virus originated from Wuhan, China, and was spread all over the world, representing no RNA variations. Measurement of antibodies to SARS-CoV-2 will improve disease management if used correctly. In late 2019, China reported a cluster of atypical pneumonia causative agents, responsible for the severe acute respiratory syndrome (SARS), rapidly spreading across the globe through human interactions and respiratory transmissions and caused the pandemic, Sequencing and searching the presence of IS elements in Covid-19 ss RNA seems to be essential to understand pandemic and the delayed process of vaccination. 8 copies of IS1 and another number of copies of IS2, and IS3 were identified in chromosomes and plasmid of Escherichia coli and other enteric infectious, water, soil, and environment born. IS was also observed in plasmids of hybrid E. coli –K-12. It is obvious to the author that observing the changing copies of IS significantly influenced the infective modes of bacteria, viruses, and their evolution [10,14], compared to wild types that could have
been widely used for the diagnosis of acute (current) SARS-CoV-2 infections [16,17]. To study and compare the infective mode of corona and other SARS and MERS, H1N1, viruses, it is good that parallel to their phenotypical SEM (Scanning Electron Microscopy) views, molecular biology of those viruses need to be studied by IS1 insertion sequence among all possible corona mutants, their wild types, among pneumococcus and other related microbes, that predominate during infections and in increasing fatality. After an incubation period of 3 days, coronavirus can cause the symptoms of a common cold, including nasal obstruction, sneezing, runny nose, secretions, and coughing [16-18].
Figure2A
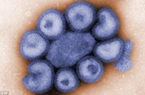
Figure2B
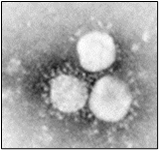
Figure2C
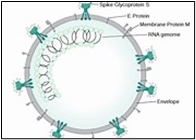
Figure2C-1
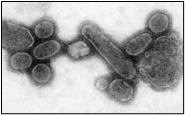
Figure2D
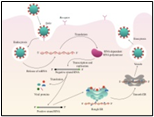
Figure2E
Figure. 2A shows the SEM (Scanning Electron Microscope) views of H1N1, (B) Influenza virus colony, (C) SERS-COV-2 virus aggregating by their spike proteins in colony formation. The molecular structure of -COV-2 RNA-virus, ss RNA, (2C1) transcribe coat- spike (S) glycoprotein, enzymatically initiate ACE-I, II (angiotensin-converting enzyme, I and II) surface activities of the host cell, [19,20] to adhere membrane of lung cells. OC43 haemagglutinin-type virions are released from host cells after rupturing the respiratory cell membrane. New virions transmit the infection via airborne droplets to the healthy recipients. The virus replicates/multiplies spontaneously in support of ss m-RNA and r-RNA of respiratory lung cells. (D) shows the outbreak of Spanish flu in the 1918 pandemic. Viral clusters and molecular biology significantly represent the similarities of their spike proteins and colony formations. The envelope glycoprotein is the characteristic feature, the chemistry of adherence. By replicating and synthesizing spike protein. (E) shows the proposed cycle of m-RNA ss RNA Covid-19, which directly is involved in the rapid growth of spike (S) proteins, capsid formation, and multiplying virus particles in lung cells. 3’ ss m-RNA of Covid-19, recognizes 5’r-RNA of infected cells, translates glycoprotein, envelope membrane matrix protein, and new coronavirus particles, to spread infections.
Coronaviruses as reported found in avian and mammalian species, resemble morphologically phenotypically similar, but differ by their antigenic structures. Even coronaviruses of humans and cattle are found antigenically similar, varied by their pathogenic expression. As common colds symptom, respiratory infections, and coronaviruses, invade many different tissues and initiate MOF (multiorgan failure) and fatal deaths of corona patients.
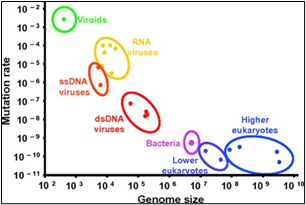
Figure 3: Genome size and mutation rate relationship
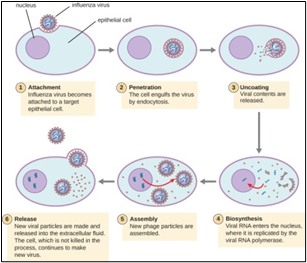
Figure 4: Tropism of Influenza Viruses on human cells
There are three types of RNA genome: dsRNA, positive (+) single-strand (+ssRNA) or negative (−) single-strand RNA (−ssRNA). If a virus has a +ssRNA genome, it can be translated directly to make viral proteins. Viral genomic +ssRNA acts like cellular mRNA. However, if a virus contains a −ss-RNA genome, the host ribosomes cannot translate it until the −ssRNA is replicated into +ssRNA by viral RNA-dependent RNA polymerase (RdRP) (see Figure 5). The RdRP is brought in by the virus and can be used to make +ssRNA from the original −ssRNA genome. The RdRP is also an important enzyme for the replication of dsRNA viruses because it uses the negative strand of the double-stranded genome as a template to create +ssRNA. The newly synthesized +ss-RNA copies can then be translated by cellular ribosomes.
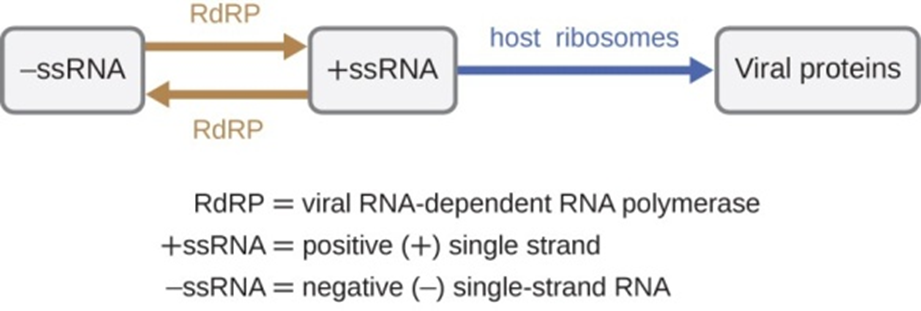
Figure 5: RNA viruses can contain +ssRNA that can be directly read by the ribosomes to synthesize viral proteins. Viruses containing−ssRNA must first use the −ssRNA as a template for the synthesis of +ssRNA before viral proteins can be synthesized.
An alternative mechanism for viral nucleic acid synthesis is observed in the retroviruses, which are +ssRNA viruses (see Figure 6). Single-stranded RNA viruses such as HIV carry a special enzyme called reverse transcriptase within the capsid that synthesizes a complementary ssDNA (cDNA) copy using the +ssRNA genome as a template. The ssDNA is then made into dsDNA, which can integrate into the host chromosome and become a permanent part of the host. The integrated viral genome is called a provirus. The virus now can remain in the host for a long time to establish a chronic infection. The provirus stage is similar to the prophage stage in a bacterial infection during the lysogenic cycle. However, unlike prophage, the provirus does not undergo excision after splicing into the genome.
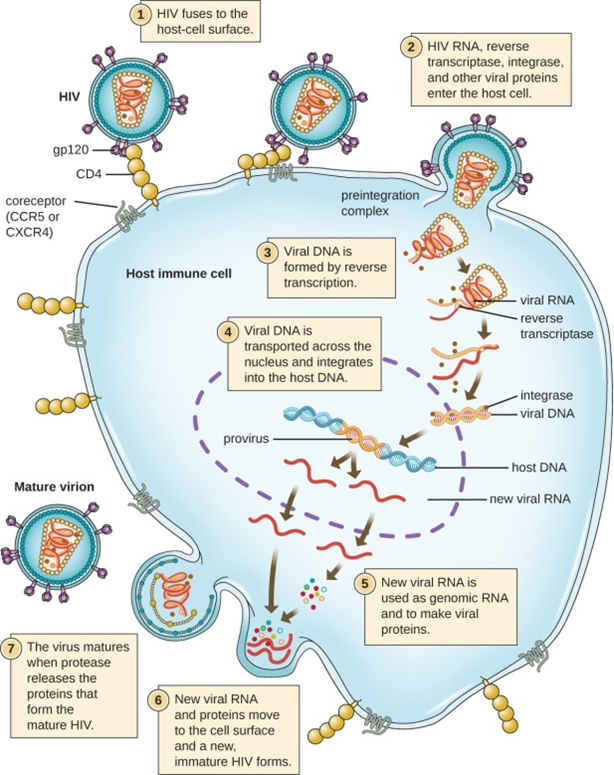
Figure 6: Represents HIV, an enveloped, icosahedral retrovirus, that attaches to a cell surface receptor of an immune cell and fuses with the cell membrane. Viral contents are released into the cell, where viral enzymes convert the single-stranded RNA genome into DNA and incorporate it into the host genome. (credit: modification of work by NIAID, NIH).
Most human coronaviruses fall under two serotypes, OC43-and 229E- surface antigen protein expressions, [1]. Fig.3, shows the genomic length and the frequency of mutation. Viruses remain at the top with 10^-2. The new strain has a change to its spike proteins -- the regions of its outer shell that dock on our cells and infect them. The change makes it a much more efficient predator. It passes quickly from cell to cell in our bodies, copying itself at a furious pace. Baric’s experiments help to explain why the 614G strain, which first emerged in Europe in February, has quickly dominated worldwide spread [19]. He says that the virus likely jumped out of bats and discovered a brand new population of human hosts, with more than 7 billion of us on the planet to infect. None of us has any immune defenses against it, so we are prime targets. Viruses with genetic advantages that help them copy themselves faster and jump more quickly between hosts to survive and to cause a pandemic,” says Baric, who is one of the world’s foremost experts on coronaviruses. His new study is published in the journal Science. The new study backs up earlier research by a team of scientists led by Bette Korber, Ph.D., at Los Alamos National Laboratory in New Mexico. The team first noticed the rapid spread of the new strain and questioned whether the virus wasn’t evolving to become more easily passed between people. Nov. 13, 2020 -- The virus that causes COVID-19 is not the same strain of coronavirus as it first emerged in China, affected and killed millions of innocents globally by corona A new study shows it has changed slightly in a way that makes it more contagious to humans., Compared to the original strain, people infected with the new strain -- called 614G -- have higher viral loads in their nose and throat, though they don’t seem to get any sicker. But they are much more contagious to others.
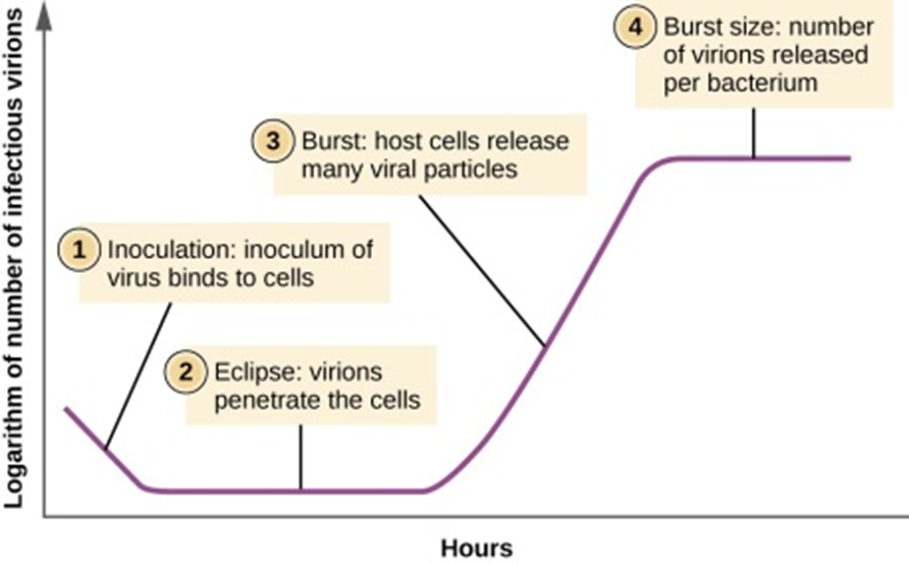
Figure 7: The one-step multiplication curve for a bacteriophage population follows three steps: 1) inoculation, during which the virions attach to host cells; 2) eclipse, during which entry of the viral genome occurs; and 3) burst, when sufficient numbers of new virions are produced and emerge from the host cell. The burst size is the maximum number of virions produced per bacterium.
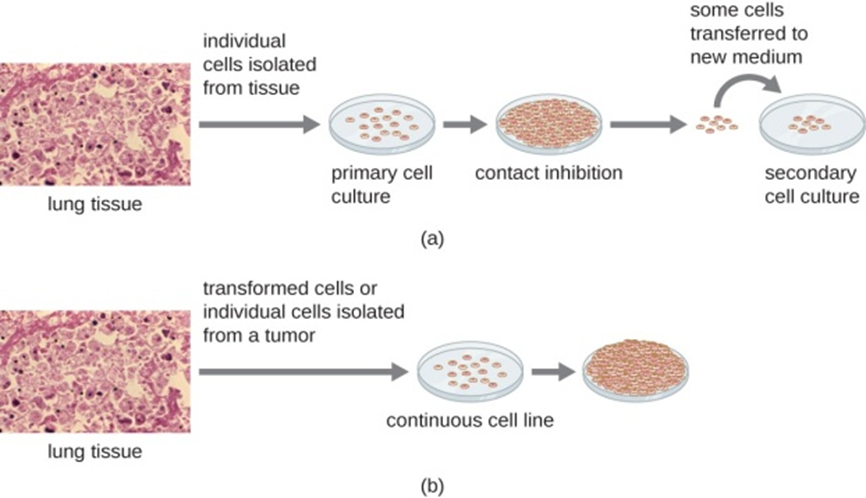
Figure 8: Cells for culture are prepared by separating them from their tissue matrix. (a) Primary cell cultures grow attached to the surface of the culture container. Contact inhibition slows the growth of the cells once they become too dense and begin touching each other. At this point, growth can only be sustained by making a secondary culture. (b) Continuous cell cultures are not affected by contact inhibition. They continue to grow regardless of cell density. (credit “micrographs”: modification of work by Centers for Disease Control and Prevention)
Agglutination Assays.
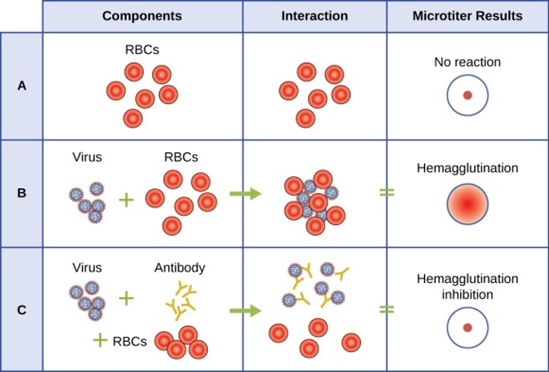
Figure 9: This chart shows the possible outcomes of a hemagglutination test. Row A: Erythrocytes do not bind together and will sink to the bottom of the well plate; this becomes visible as a red dot in the center of the well. Row B: Many viruses have hemagglutinins that cause agglutination of erythrocytes; the resulting hemagglutination forms a lattice structure that results in red color throughout the well. Row C: Virus-specific antibodies, the viruses, and the erythrocytes are added to the well plate. The virus-specific antibodies inhibit agglutination, as can be seen as a red dot at the bottom of the well. (credit: modification of work by Centers for Disease Control and Prevention)
Conclusion
Microbes and climate change are proportionally related oscopic An outbreak of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) was first reported in Wuhan City, China, in December 2019. Since then, the outbreak has grown into a global pandemic, and neither a vaccine nor a treatment for the disease, termed coronavirus disease 2019 (COVID-19), is currently available. The slow translational progress in the field of research suggests that a large number of studies are urgently required. In this context, this review explores the impact of bacteriophages on SARS-CoV-2, especially concerning phage therapy (PT). Bacteriophages are viruses that infect and kill bacterial cells. Several studies have confirmed that in addition to their antibacterial abilities, bacteriophages also show antiviral and antifungal properties.
River Ganges in India, for centuries, has been revered for its “self-cleansing and special healing properties”. More than 450 million people depend on the waters of the Ganges for many aspects of their life. In 1896, one of the first published works on Ganges water by Ernst Hankin, a British bacteriologist demonstrated the antibacterial property of Ganges water against Vibrio cholera [1]. Further work by French microbiologist D’Herelles at the beginning of the twentieth century established that the antibacterial property of Ganges water be due to a factor, later named “bacteriophage” [2] Virus Bacteria, (Unorganised without nucleus Fungi, Mould and Algae (Organize with a nucleus) are propagating and changing their non-pathogenic mode to pathogenic mode quickly and slowly based on their pro (uunorganized karyotic and Eu (organized, nuclear material). The somatic, reproductive, and immune-responsive cells of our body belong mainly to eukaryotic cells and tissues. Viral cells as prokaryotes carry ss, (single-stranded), or ds (double-stranded) RNA or DNA and transposons. Viruses can only survive if a host for such viruses is found, [14]. For survival, viruses adapt environment spontaneously by mutations and participate in evolution. Bacteria and viruses, follow their metabolic activities, either in the form of symbiosis, partial full parasitic nature, “or host–vs -graft” immune responses (IR). Bacteria the size, of 1 micron (μ) to 10 micron (μ) irrespective to host, have their own metabolic system. On the other, viruses are fully host-specific, changing from non-pathogenic to pathogenic, and lytic to lysogenic phases. Based on the signal received from hosts, bacteriophages in the case of bacteria and retro adenoviruses use animals and humans to propagate by means of infections. As per host and environment, they colonize
and grow in normal and extreme, acidophilic (pH1-2), mesospheric, and thermophile environments. Lambda, Psiex, and T4 viruses share their existence by transforming DNA to RNA among adeno, retro, and rhino, animal viruses
(human, animal, bird, and insect) to sustain host-specific activities, [14,15]. The origin of the river Ganges is known as Gomukh. Geological studies have proven that the Himalayas had emerged at a site where the Tethys Ocean once existed, as a result of collision between the Indian and Eurasian tectonic plates. Hence, Himalayas’ marine origin is known and it has been intriguing to find marine fossils high in the Himalayas [3].
References
- Fedoroff, Nina V. "McClintock’s challenge in the 21st century." Proceedings of the National Academy of Sciences 109, no. 50 (2012): 20200-20203.
- Starlinger, Peter, and Heinz Saedler. "IS-elements in microorganisms." In Current Topics in Microbiology and Immunology/ Ergebnisse der Microbiologie und Immunitätsforschung, pp. 111-152. Berlin, Heidelberg: Springer Berlin Heidelberg, 1976.
- Saedler, H., D. Ghosal, and P. Nevers. "The role of gene rearrangement in evolution." In Modern Trends in Human Leukemia III: Newest Results in Clinical and Biological Research, pp. 507-511. Berlin, Heidelberg: Springer Berlin Heidelberg, 1979.
- Babakhani, Sajad, and Mana Oloomi. "Transposons: the agents of antibiotic resistance in bacteria." Journal of basic microbiology 58, no. 11 (2018): 905-917.
- Ruediger Schmidt (1982),Transposon; Springende Gene in Bakterien, Erbforschung Heute. Herousgegeben vom Walter Klingmueller, Verlage Chemie, Basel, Switzerland.
- Gu, Hongjing, Qi Chen, Guan Yang, Lei He, Hang Fan, Yong- Qiang Deng, Yanxiao Wang et al. "Adaptation of SARS-CoV-2 in BALB/c mice for testing vaccine efficacy." Science 369, no. 6511 (2020): 1603-1607.
- Shapiro, James A. "Molecular model for the transposition and replication of bacteriophage Mu and other transposable elements." Proceedings of the National Academy of Sciences 76, no. 4 (1979): 1933-1937.
- Bukhari, A. I., and D. Zipser. "Random insertion of Mu-1 DNA within a single gene." Nature New Biology 236, no. 69 (1972): 240-243.
- Georgiou, Ioannis, Dimitrios Noutsopoulos, Eftychia Dimitriadou, Georgios Markopoulos, Anastasia Apergi, Leandros Lazaros, Terpsi Vaxevanoglou, Kostas Pantos, Maria Syrrou, and Theodore Tzavaras. "Retrotransposon RNA expression and evidence for retrotransposition events in human oocytes." Human molecular genetics 18, no. 7 (2009): 1221-1228.
- Brahma, Nitosh, Arne Schumacher, John Cullum, and Heinz Saedler. "Distribution of the Escherichia coli K12 insertion sequences IS1, IS2 and IS3 among other bacterial species." Microbiology 128, no. 10 (1982): 2229-2234.
- Brahma, N. K. (1999). Enterobacterial 026: MRHA-plasmid and its possible Genetic Transformation into Auxotrophic Escherichia coli K-12 C600 NS. Indian J Microbial, 38, 43-50.
- Brahma, N. K. (1999). Genetically Engineered Hybrid 5405 E. coli K-12 fimbriae in vaccination With Balb/c mice against 026: serotype EPEC diarrhea. Ind. J. Chem. Engg, 39(1).
- Brahma NK (2020) Concept for Possible Development of Vaccine against Corona in Escherichia coli K-12, Yale C600 Strain, EC Clinical and Medical Case Reports 3.6 (2020): 111-116.
- https://www.onlinescientificresearch.com/articles/understanding- rapid-transmitting-invasive-corona-pandeics-through-aair- istnis- crisprcas9-dna-editing.pdf
- Greaney, Allison J., Tyler N. Starr, Pavlo Gilchuk, Seth J. Zost, Elad Binshtein, Andrea N. Loes, Sarah K. Hilton et al. "Complete mapping of mutations to the SARS-CoV-2 spike receptor- binding domain that escape antibody recognition." Cell host & microbe 29, no. 1 (2021): 44-57.
- Natural Bacteriophages' Potential—A Direct Weapon Against Bacteria[16]Natallie Crowley, 2020, A Look Into the Epigenetics of a Coronavirus Infection Epigenetics, March 10,Diseases & Disorders, Educationally Entertaining
- Greaney, Allison J., Tyler N. Starr, Pavlo Gilchuk, Seth J. Zost, Elad Binshtein, Andrea N. Loes, Sarah K. Hilton et al. "Complete mapping of mutations to the SARS-CoV-2 spike receptor- binding domain that escape antibody recognition." Cell host & microbe 29, no. 1 (2021): 44-57.
- Julia Evangelou Strait (2020). Jumping genes' help stabilize DNA folding DNA folding patterns, Washington University School of Medicine, 24 January 2020, Phyorg.
- Chymen Chemicals(2021) News and Announcement, Concerning Variants of SARS- CoV-2,Featured Article from 2021- 01-25.
- Bernstein, Kenneth E., Zakir Khan, Jorge F. Giani, Duo-Yao Cao, Ellen A. Bernstein, and Xiao Z. Shen. "Angiotensin-converting enzyme in innate and adaptive immunity." Nature Reviews Nephrology 14, no. 5 (2018): 325-336.
- Southern, Edwin Mellor. "Detection of specific sequences among DNA fragments separated by gel electrophoresis." J mol biol 98, no. 3 (1975): 503-517.
- Jaidip Dogra, Livodip Dogra, 2020, “Opportunistic Chlamydia in Corona, Jaipur Central Govt Health Scheme, ”Pratidin Bengali newspaper, 25th May, p:1 and5
- Braude AI, Davis, CE, Fierer j, (1981) Medical Microbiology and Infectious Diseases, Saunders Pub, International Text Book of Medicine.Page:619, D.A.J. Tyrrell Corona Virus.
- David A.J. Tyrrell and Steven H. Myint, (1980) Medical Microbiology. 4th edition,
- Cui, Jie, Fang Li, and Zheng-Li Shi. "Origin and evolution of pathogenic coronaviruses." Nature reviews microbiology 17, no. 3 (2019): 181-192.
- Ahn, Dae-Gyun, Wooseong Lee, Jin-Kyu Choi, Seong-Jun Kim, Ewan P. Plant, Fernando Almazán, Deborah R. Taylor, Luis Enjuanes, and Jong-Won Oh. "Interference of ribosomal frameshifting by antisense peptide nucleic acids suppresses SARS coronavirus replication." Antiviral research 91, no. 1 (2011): 1-10.
- Broecker, Felix, and Karin Moelling. "Evolution of immune systems from viruses and transposable elements." Frontiers in microbiology 10 (2019): 51.
- Leslie, Mitch. "T cells found in coronavirus patients ‘bode well’for long-term immunity." (2020): 809-810.
- Crispr/Cas-9 and RAG ½ T-cell diversity; Stanford Bioengineering Schools of Engineering & Medicine, .. "Development of CRISPR as a prophylactic strategy to combat novel coronavirus and influenza"By Brenda Goodman, MA
- Brahma NK (2020) Nano-Engineering, Vaccination & Diagnosis in Combating Corona : The Recent Trends”, 25.11.2020, ODTW (Oneday Technical Webinar), CHDB, by WBSC, The Institution of Engineers, India,” Feasibility Searched for Nanocoating on lung alveoli cells”. COVID-19: Bacteriophage Could Decrease Mortalityhttps://www.liebertpub.com › doi › phage.2020.0014
- -Jun-2020 — The pandemic of the coronavirus disease (Covid-19) has caused the death of at least 270,000 people as of the 8th of May 2020.
- Ganga Water and Phage.


